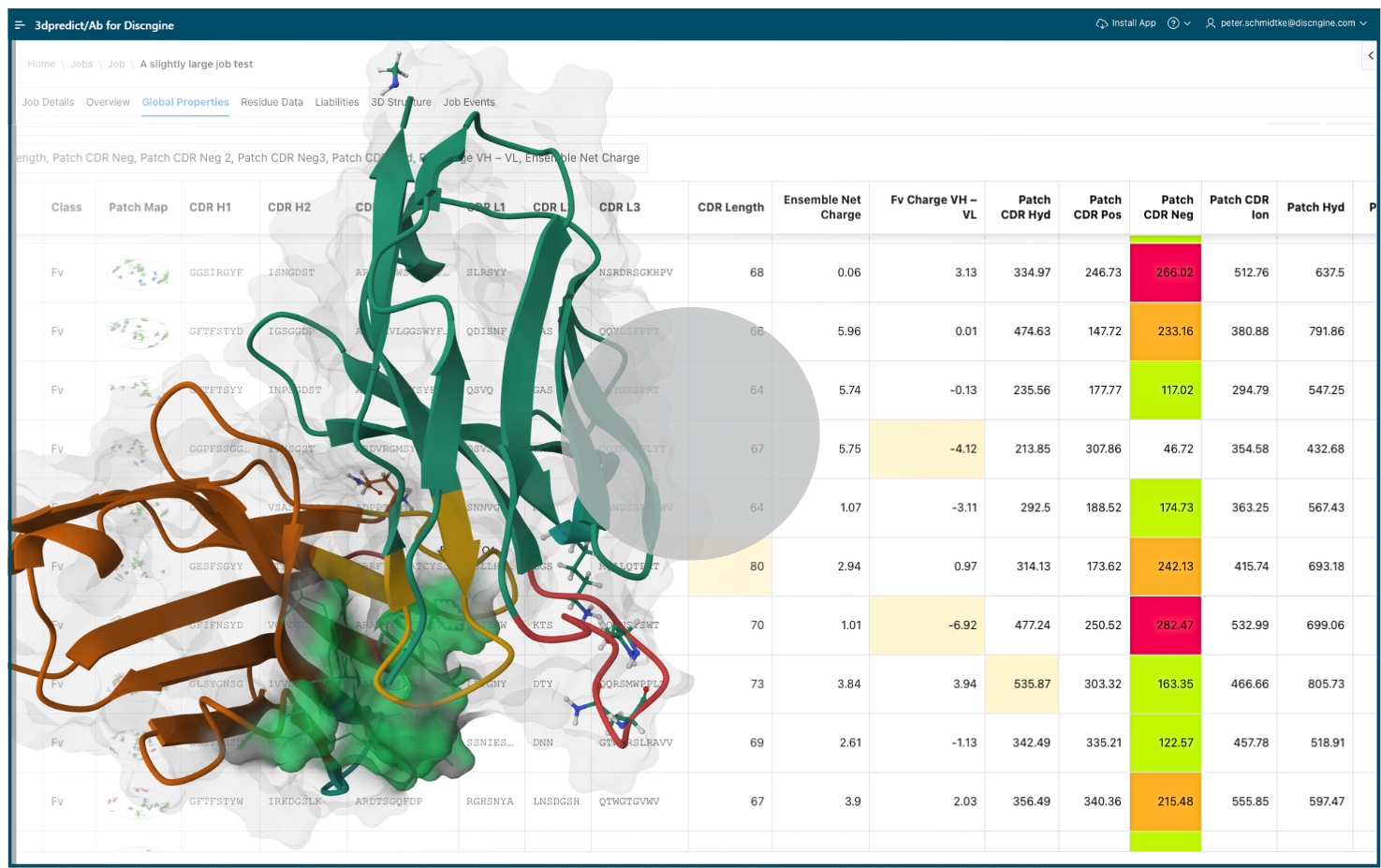
Screen and rank antibody candidates for developability & liability with ease
3dpredict/Ab is a Software-as-a-Service (SaaS) platform providing ensemble-based predictions to help overcome the single-structure bias, supporting quick candidate selection and progression through the discovery pipeline.
It allows antibody modelers and engineers to identify potential liabilities and compare antibody variants based on physics-based property calculations.
Its scalable cloud architecture supports high-throughput screening and can be integrated with existing discovery workflows for efficient data management and analysis.
Successfully prioritize antibodies to accelerate your lead optimization phase

Ensemble-based predictions at your fingertips
Accurate antibody developability insights from ensemble-based protein property calculations accounting for pH-dependence and flexibility

User-friendly platform built for efficient discoveries
Intuitive web-based interface designed for both small and big antibody discovery groups in pharma and biotech

Instant access to focus on science, not IT hassle
Immediately accessible through a highly reliable and secure cloud environment (ISO 27001) without any deployment efforts
Streamline your antibody discovery workflows with accurate ensemble-based predictions
Sequence/structure upload & calculations setup
Submit antibody sequences or structures through the UI or API, with multiple input file types (FASTA, XLSX, PDB).
Explore several antibody formats, including Fv, scFv, VHH, Fab, bispecifics, ADCs, and custom constructs.
Set up calculations for conformational and pH sampling.
Structural ensemble generation & property calculations
Generate structural ensembles using stochastic titration and accelerated grand canonical sampling to capture conformational and protonation-state variability.
Compute 100+ whole protein and residue-level properties/descriptors to assess antibody aggregation, viscosity, solubility, charge distribution, and other developability-related outcomes.
Simulate complex molecular systems, including non-natural amino acids and conjugated systems.
Avoid static conformation bias and thresholding effects.
Get reproducible results with confidence intervals.
Developability & liability screening
Leverage a high-performing proprietary algorithm based on four determined rules for therapeutic antibody profiling against clinical-stage antibodies (Thorsteinson et al; mAbs 13 (2023) 219-235).
Perform large-scale, high-throughput screening to rank and prioritize antibody candidates early in discovery.
Automatically detect liability hot spots, including aggregation-prone regions, and PTMs.
Visualize surface features through standardized 2D protein patch maps.
Analysis & prioritization
Enhance the predictivity of your in-house AI models with high-quality data - boost your HIC retention time predictions vs static models.
Leverage sequence visualizer to assess annotated liabilities.
Identify germline sequences for potential back-mutations and compare residue-level properties across candidates.
Map detected liabilities back to the 3D structure to decide on the next optimization move.
Identify outliers within your reference dataset using customizable coloring rules and multiple plotting options for clear visual comparison.
What’s new
3dpredict/Ab releases: Discover major updates for improving antibody developability property predictions
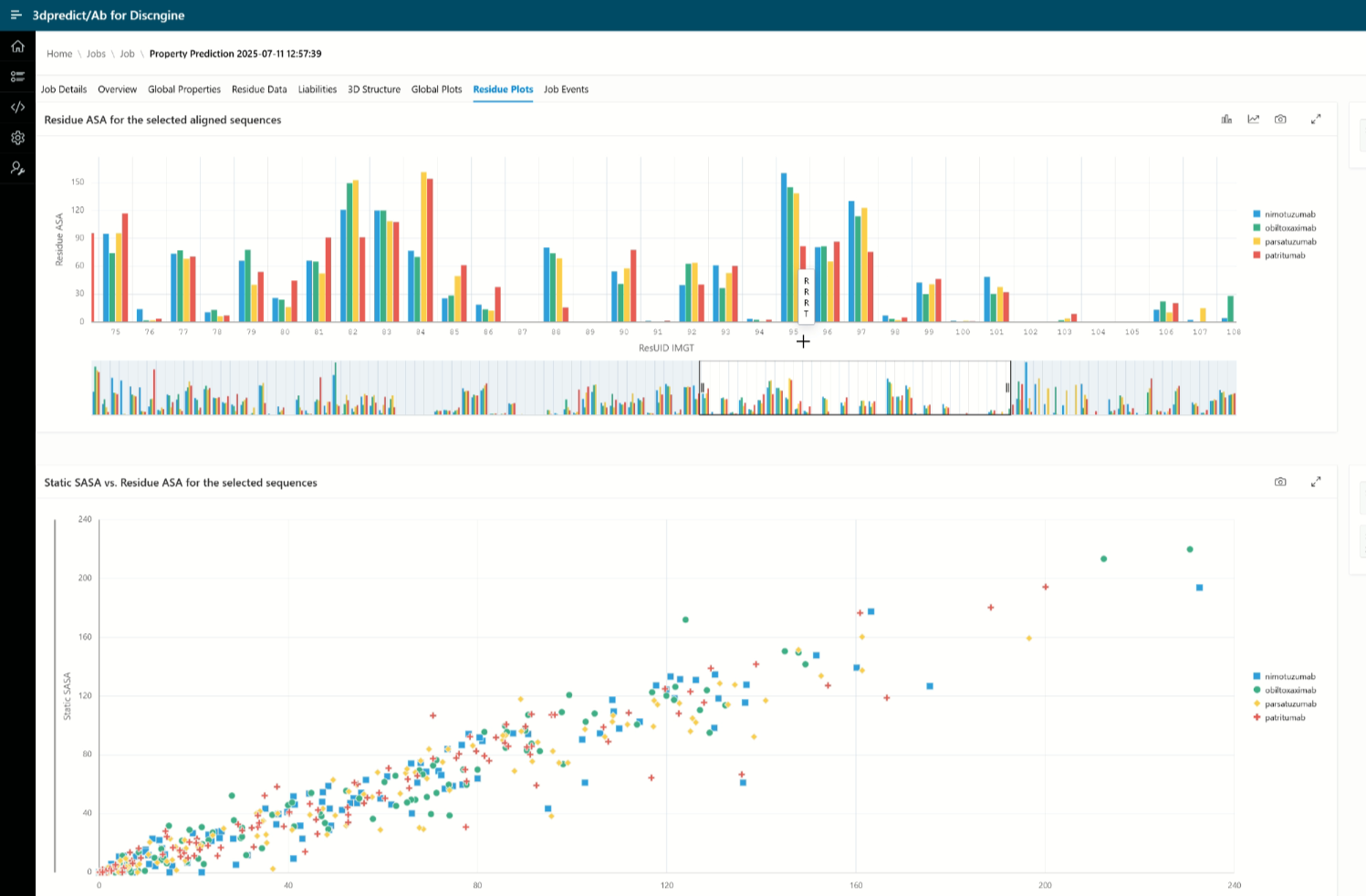
3dpredict/Ab 2025.09
This new version allows users to build custom reference collections, calculate properties of non-standard antibody formats (such as ADCs), and explore multiple new property plots for enhanced visualization and analysis.
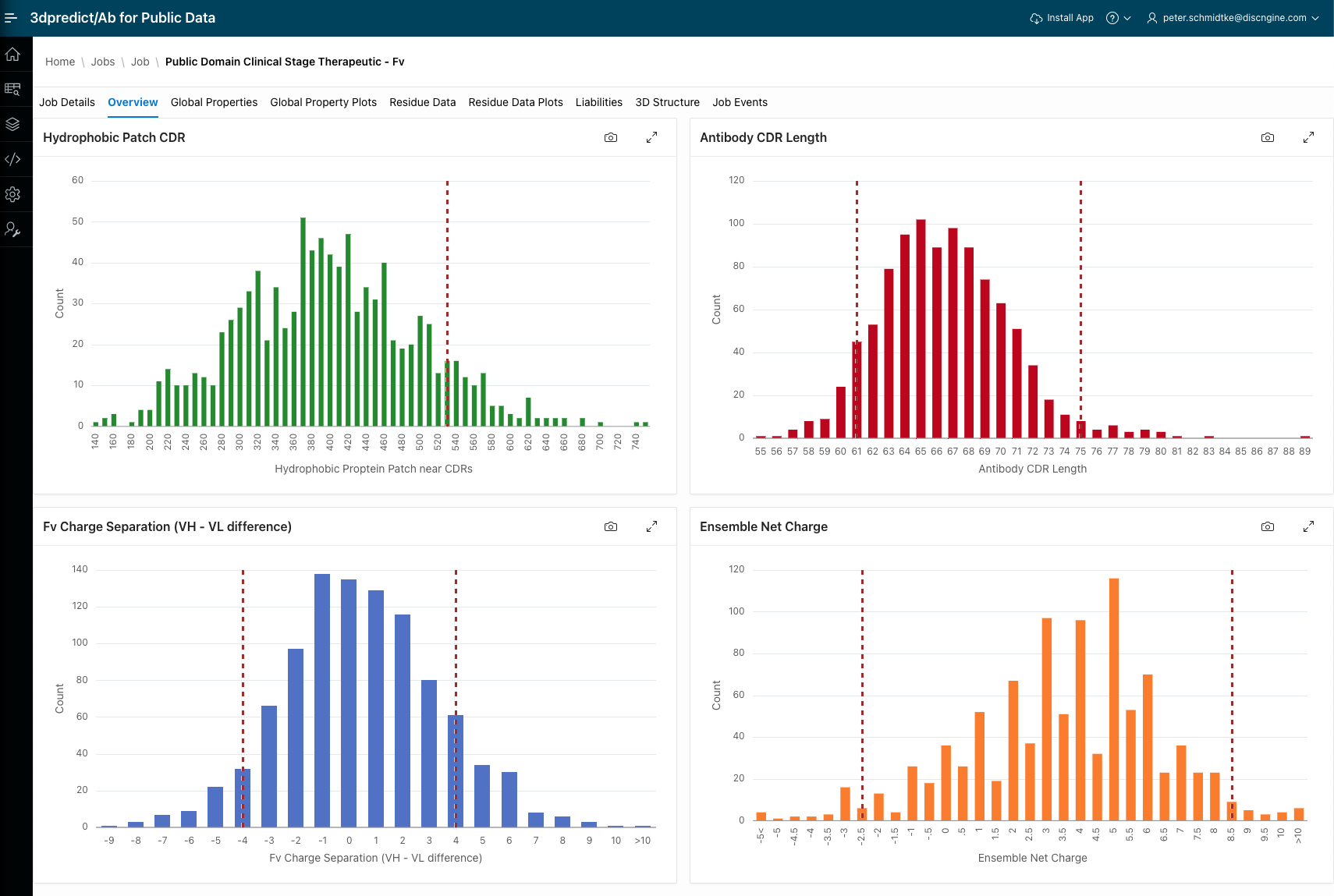
3dpredict/Ab 2025.12
This release adds a curated set of 1000 clinical-stage antibodies with predicted properties, enabling discovery teams to compare new candidates against them and make more informed developability decisions.
For technical documentation and feature tutorials, please visit the public documentation page.


High throughput
3dpredict/Ab can process large antibody sets in parallel, performing up to 30,000 ensemble calculations per day to support early identification of candidates with favorable properties and reduced liabilities.

Secure cloud environment
Discngine complies with ISO 27001 standards for information security, ensuring high protection of research data.
We offer secure application and data hosting on the Discngine Cloud Infrastructure, fully managed by our experts.

Integration
The REST API allows seamless integration with your in-house IT ecosystem to complement commonly used tools with high-quality insights for decision-making.
3dpredict/Ab was developed in collaboration with Chemical Computing Group (CCG).

Frequently Asked Questions
Have more questions?
-
3dpredict/Ab is a cloud-based platform that generates conformational ensembles of antibody structures and predicts high-quality physicochemical properties at scale.
3dpredict/Ab enables antibody discovery teams to assess developability and liability risks and contributes to antibody candidate profiling, ranking and screening at different stages of antibody discovery.
-
3dpredict/Ab relies on a physics-based method for antibody 3D structure predictions and ensemble-based property predictions.
It uses stochastic titration and accelerated grand canonical sampling to generate multiple conformations and protonation states, thereby accounting for structural flexibility and pH-dependent charge effects.
Developability-relevant physicochemical properties are then computed across the structural ensemble. This helps reduce single-structure bias and enables more accurate early-stage candidate ranking.
-
3dpredict/Ab uses a proprietary therapeutic profiling approach to compare predicted antibody properties against a reference dataset comprising clinical-stage therapeutic antibodies.
The "rule of four" refers to a set of four structure-based descriptors to evaluate the developability and "drug-likeness" of monoclonal antibodies (mAbs) during early-stage drug discovery. These rules are designed to filter out candidates prone to high aggregation, viscosity, and poor pharmacokinetics (PK) by analyzing the Fv region's surface properties, such as hydrophobicity near CDR, ensemble net charge, CDR lengths, and Fv charge separation (Thorsteinson et al; mAbs 13 (2023) 219-235).
-
3dpredict/Ab is accessible via a web interface, with no local installation required, and runs in Discngine’s secure cloud environment. Discngine is ISO/IEC 27001:2022 certified, with certification covering software development, customer support, and Software-as-a-Service (SaaS) hosting of software solutions.
3dpredict/Ab provides API access for batch sequence submissions and results retrieval which enables integration into existing discovery workflows.
-
3dpredict/Ab supports a wide range of antibody and antibody-derived formats, including Fv fragment, Fab, single-chain Fv (scFv), VHH (nanobodies), bispecifics, antibody-drug conjugates (ADCs), and custom constructs. Antibody candidates can be submitted as sequence files (FASTA), structure files (PDB), or batch input tables (XLSX), enabling both single-candidate analysis and high-throughput screening.
It is also possible for us to handle more complex antibody formats. Please get in touch with us for more information.